File:World Map-DEC.jpg
From Human Embryology
Size of this preview: 800 × 446 pixels. Other resolution: 1,000 × 558 pixels.
Original file (1,000 × 558 pixels, file size: 91 KB, MIME type: image/jpeg)
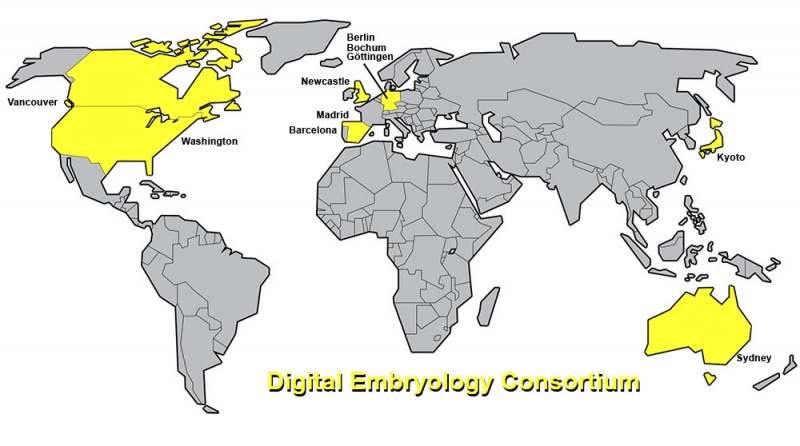
World Map - Digital Embryology Consortium
Distribution of Embryo collections shown on world map (2016).
Listed below (alphabetically) are the human embryo collections (2016) that are currently contributing to the Consortium. Dr Mark Hill, UNSW Australia established and has coordinated the consortium scanning and online content. Prof. Shigehito Yamada has agreed to be the deputy coordinator of the consortium.
| DEC OMERO Server | ||||||||||||||||
| ||||||||||||||||
- Blechschmidt Collection – University of Göttingen, Germany Prof. Christoph Viebahn (Head of Embryology Department).
- Carnegie Collection – National Museum of Health and Medicine Elizabeth Lockett (curator NMHM).
- Domenech-Mateu Collection – Universitat Autònoma de Barcelona, Prof Rosa Mirapiex.
- Hinrichsen Collection – Ruhr-University Bochum, Prof. Beate Brand-Saberi (Head of Embryology Department).
- Embryological Collection – Museum für Naturkunde, Leibniz Institute for Research on Evolution and Biodiversity. Dr Peter Giere (curator of the embryological collection).
- HDBR and HuDSeN Collection – Newcastle University, Prof. Susan Lindsay.
- Kyoto Collection – Kyoto University, Prof. Shigehito Yamada.
- Orts-Llorca Collection – Complutense University of Madrid, Institute of Embryology, Prof. Jose Francisco Rodriguez Vazquez.
- Perry-Arey-Milligan Collection – University of British Columbia, Prof. Virginia Divert.
- UNSW Collection - UNSW Australia, Dr Mark Hill
Main Page | Embryo Collections | Slide Scanning | Image Server | News | Links | Test page | Site Map
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 06:40, 12 March 2016 |  | 1,000 × 558 (91 KB) | Z8600021 (talk | contribs) |
You cannot overwrite this file.
File usage
The following 5 pages use this file: