DEC Online Database
About the Online Database
| DEC OMERO Server | ||||||||||||||||
| ||||||||||||||||
The collection scanned images are currently available as a test site with only a few sample images loaded. This is currently only in a trial, final database may vary from this draft structure following user testing.
There is no unregistered user access available to the server. All users must apply for an account and agree to abide by our access and reuse conditions before being issued a user account. Individual user accounts currently only have access to specific folders. Access to other collection folders can be managed through user rights management.
Database Images:
- can be accessed and downloaded remotely through any web browser link to DEC Database.
- can be imported, accessed and downloaded remotely through OMERO.insight or ImageJ/Fiji.
Importing Data
The two methods for importing data are shown below.
OMERO.insight
Images are added to the database by the process called "Importing Data" using the software OMERO.insight Version 5. Please read the DEC OMERO training manual before uploading images to the test version of the image server.
20 July 2017 - Please note error on Page 4 of training manual the server address to enter should be 149.171.80.223 not the full web address shown in the original manual (http://149.171.80.223:8080). My apologies for this error.
ImageJ/Fiji
Images can also be added to the database as well as adding ROI's to existing images through ImageJ/Fiji. I am currently preparing a DEC manual and video to demonstrate this process. Best place to currently begin using ImageJ/Fiji is with the original OMERO help page Using ImageJ with OMERO and the 2016 demonstration video below.
|
<html5media width="500" height="400">File:Fiji omero manual workflow 2016.mp4</html5media> |
Collections
The DEC collections have been organised into project folders named after the original collection name or location. Scanned images in the JPGXR format have been initially added to the database using OMERO.insight. Images have been added to the database as the collection owner/curator, and this also tags the image file to the original collection. Note that only admin users can add images as other users, if you log-in and add images they will appear in the database as "owned" by you.
All users can currently see all images in the database by selecting the appropriate collection and owner.
A collection then has a folder for each embryo, named as in the original collection. Within this main folder are sub-folders for each slide, as numbered in the original collection.
The main folder also has additional "Project Details" and "Tags" as shown in the sub-headings below.
Embryo
- Each embryo has own main folder (with the original embryo name as it exists in collection).
- Within this main folder each slide currently has its own sub-folder.
Project Details
- Project details are added to the embryo main folder and include.
- Original embryo name as shown on slide
- Carnegie stage (if known)
- Crown-Rump Length (CRL) in mm
- Plane of sectioning - Sagittal, Transverse, Frontal (coronal), Oblique
- Histology stain(s) used for this embryo set
- Total number of slides available (from catalogue).
Tags
These are new tags added to the online database. Some relate to whole embryo and some are for individual slides.
Currently "Tags" cannot be moved between collection groups. In order to add the entire Tab Set and Tag list has to be added individually to each collection.
The following subheadings are the "Tag Set"folder.
Collection
- Blechschmidt - (whole embryo) Should act as Gottingen collection identifier tag when searching whole library.
- Hinrichsen - (whole embryo) Should act as Bochum collection identifier tag when searching whole library.
Plane
- Sagittal - (whole embryo) plane dividing the body into left and right halves.
- Transverse - (whole embryo)
- Frontal - (whole embryo) Coronal planes in collection are also listed with this tag.
- Oblique - not an exact anatomical plane. May include more details in Project Details section.
Histology
Note - Some embryos have 2 to 3 different stains used in the serial sections. Too difficult at this stage to tag each slide so currently (whole embryo) may be tagged with multiple stains.
- Toluidine Blue - (whole embryo) Histology stain, nucleus blue and cytoplasm light blue. (Example - Hindrichsen ME65 heart)
Housekeeping
Related to database activities. Should allow easy identification of upload fails.
- Fail - (individual slide) added to individual slides that fail to upload with OMERO.insight (Example - Hindrichsen Embryo 283 slides 478-484, Embryo 335 slides 160-164)
Sample Images
July
Locally uploaded image samples.
- Blechschmidt Collection in Christop Viebahn folder http://149.171.80.223:8080/webclient/?show=dataset-641
- Hinrichsen Collection in Beate Brand-Saberi folder http://149.171.80.223:8080/webclient/?show=dataset-735
- Hubrecht Collection in Peter Giere folder http://149.171.80.223:8080/webclient/?show=dataset-786
June
Selected slides form the Blechschmidt Collection are available for registered user access. Please advise me if you are unable to see the linked files below following log-in.
- 2.57 mm sagittal - 22 slides total 23 (1954-08-10)
- 3.4 mm sagittal - 82 slides total 87 (1954-04-09)
- 4 mm sagittal - 3 slides total 4 (1951-05-05)
- 5.8 mm frontal - 54 slides total 77 (1954-09-20)
- 6.5 mm sagittal - 40 slides total 40 (1953-09-29)
- 11 mm frontal - 29 slides total 76 (1951-09-01)
2016 Test Image Link
Note - You will need to login to access the original image through the links, images appearing on this current page are only screenshots.
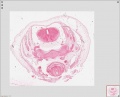
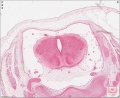
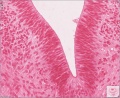
Sydney folder -> 2010-08-29-1 -> Stage 22 Human Embryo (Mark Hill-001.svs) dimensions 32538 x 28770
Image Formats
The original decision was that scanned slides would be saved in JPGXR format as these are substantially smaller than the uncompressed format. Note that OMERO server will accept embryology images in a variety of formats, including uncompressed images, the problem will be the overall size of the database would be unmanageable.
Following collection scanning, each collection manager should also hold an uncompressed archive of their own images. Should you require images of this quality/resolution, please contact the original collection managers.
CZI
The Axioskan slide scanner generates images in the CZI file format that was developed by ZEISS to specifically meet the requirements of imaging in microscopy. This is an excellent uncompressed research image format, but generates images that would be far too large for easy access on the DEC image database.
JPG XR
The Axioskan slide scanner software can also save or convert images into the JPG XR format. Note that the image file extension will still be shown as CZI, but this will be a "compressed" file. This format was selected as it has not licensing requirements, generates smaller good quality images, and the scanner can readily batch generate this format.
The JPG XR file format is a still-image compression standard and format for continuous tone photographic images, based on technology originally developed and patented by Microsoft under the name HD Photo (formerly Windows Media Photo). It supports both lossy and lossless compression, and is the preferred image format for Ecma-388 Open XML Paper Specification documents (Wikipedia). This file format is also an International Organization for Standardization ISO/IEC TR 29199-1:2011.
There are several research conversion programs, including Fiji (ImageJ), that can both read and generate this image format.
Additional non-research programs are available for image conversion into this form (Google search JPGXR conversion).
- Caution - Users should be extremely careful when downloading software from the internet and should contact their local IT managers and use anti-virus software to screen before opening, installing or running any such downloaded program.
JPEG 2000
OMERO Bioformats should also be compatible with the [ JPEG200 image format, though we have had some initial issues with this on the database. For example the image will upload to the database, but will not be able to generate the thumbnail or process the required pyramid.
- Problem - One Image 3 formats only the JPG version has generated a thumbnail image, both the TIF and J2K versions show that they are still "generating a pyramid".
- Problem - J2K image similar problem.
- OK - JPF image photograph not a scanner image uploaded and displayed without a problem. (see JPF extension)
JPEG 2000 is both a codec and a file format and the standard is in many parts.
- Part 1 giving (mostly) codec information (how to compress/decompress image data), with a container file format annex (JP2).
- Part 2 gives many extensions, and a more comprehensive container format (JPX).
- Zeiss JPEG-XR cannot be ported to the Olympus files without major work.
Apply for DEC Account
For user account requests, please contact Dr Mark Hill.
Terms of Use
By using the DEC OMERO Server, you agree to the following:
- You agree to the specific conditions set by each collections terms of image reuse, publication and copyright.
- Reuse of images from multiple collections requires approval from each collection.
- You use this service at your own risk, and we provide NO GUARANTEE OR WARRANTY WHATSOEVER regarding uptime, security, data provenance or access, or any other service.
- You should not use the DEC OMERO Server as a repository for any important data, or data that must be held in any secure or specific way.
- The responsibility for any data loaded onto the DEC OMERO Server is entirely your own.
- We reserve the right to remove any images, data, accounts or other data or structure without notice or request.
- We welcome comments or ideas from your use of and experience with the DEC OMERO Server.
About OMERO
OMERO is client-server software for visualization, management and analysis of biological microscope images. © 2000-2016 University of Dundee & Open Microscopy Environment. Creative Commons Attribution 4.0 International License OME source code is available under the GNU General public license or through commercial license from Glencoe Software.
Version - OMERO.web 5.0.8-ice35.
References
<pubmed>20513764</pubmed>
<pubmed>22373911</pubmed>
Links
- The Open Microscopy Environment
- OMERO 5.4 Download
- OME Help
- Using OMERO.web
- Training Course Manual
- Importing Data with OMERO.insight Version 5
- JCB DataViewer
- Fiji (ImageJ)
- Image formats
- Photoshop JPEGXR Plug-in (from Microsoft)
Main Page | Embryo Collections | Slide Scanning | Image Server | News | Links | Test page | Site Map